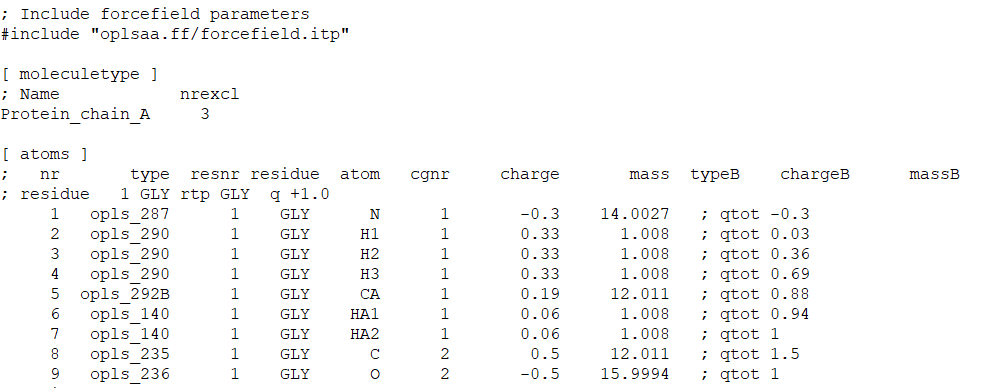
Tutorial: Modelling post-translational modified proteins with GROMACS – Dr Anthony Nash MRSC: computational chemistry and life sciences

RTP File Extension | Gromacs Residue Topology Parameter File | Associated Programs | Free Online Tools - FileProInfo

TopoGromacs: Automated Topology Conversion from CHARMM to GROMACS within VMD. - Abstract - Europe PMC













![PDF] GROMACS: A message-passing parallel molecular dynamics implementation | Semantic Scholar PDF] GROMACS: A message-passing parallel molecular dynamics implementation | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/625ddfe164e3a25405672ecc1ca17d05353ff182/8-Table1-1.png)